Chua, Physilia . Metabarcoding for hope: A large-scale molecular dietary study of the western capercailles (Tetrao Urogallus). Given at the BRC16/19 SAMBA meet 2021 - Biodiversity monitoring and mapping (hosted by UFPA, Belem, Brazil). 10 September 2021.
Presentations
Ariza, Maria. Calibrating soil eDNA vegetation assessments: a blast from the past Given at the BRC16/19 SAMBA meet 2021 - Biodiversity monitoring and mapping (hosted by UFPA, Belem, Brazil). 08 September 2021.
Moiloa, Ntwai. Species-tree diffusion models and their applications in campion (Silene) phylogeography. Given at the Plant ID Seminar Series Molecular identification of plants for science and society, Series 2: Modern tools and old methods: Descriptive taxonomy and phylogeny (hosted by NHM, UiO, Oslo, Norway). 25 March 2021.
Quatela, Anne-Sophie. Polyploid species, the Achilles’ heel of molecular taxonomy? A study case of (sub)arctic Silene sect. Physolychnis. Given at the Plant ID Seminar Series Molecular identification of plants for science and society, Series 2: Modern tools and old methods: Descriptive taxonomy and phylogeny (hosted by NHM, UiO, Oslo, Norway). 25 March 2021.
Michel, T. 2021. The Victorian parlour plant and the survival of the unfit. Given at the Plant ID Seminar Series Molecular identification of plants for science and society, Series 1: Biodiversity time travels: The use of historical plant genomes in biodiversity research (hosted by NHM, UiO, Oslo, Norway). 25 February 2021
Woudstra, Y. 2020. Obtaining small loci from large genomes: Customised nuclear target enrichment for estimating phylogenetic relationships in Aloe (Asphodelaceae). Given at the ForBio Annual Conference. Digital Conference (hosted by NHM, UiO, Oslo, Norway). 04 December 2020
Polling M., 2020. Image Recognition for Improved Pollen Identifications. Virtual Presentation. Part of AI4BIO hosted at Naturalis Biodiversity Center, Leiden, The Netherlands. 02 November 2020.
Woudstra Y, 2020. Molecular identification and phylogenomic resolution of Aloe vera and relatives using customised nuclear target enrichment. At: Botany 2020. Virtual. 27-31 July 2020.
Quatela A-S, 2020. Exploring the utility and limits of target enrichment methods to study polyploidy and reticulate evolution. At: Botany 2020. Virtual. 27-31 July 2020.
Moiloa N, 2020. On the phylogenetic position and biogeographic origins of southern African Silene. At: University of Gothenburg. Gothenburg, Sweden. June 2020.
Michel T, 2020. Ancient DNA, mapDamage, and the Paleomix pipeline. At: Norwegian University of Science and Technology. Trondheim, Norway. May 2020.
Ariza M, 2020. Molecular plant identification for biodiversity assessment. At: CEES, University of Oslo. Oslo, Norway. 10 March 2020.
Woudstra Y, 2020. New DNA barcodes for Aloe derived from NGS data. At: Royal Botanic Gardens, Kew. London, UK. 3 February 2020.
Woudstra Y, 2020. GGBC Taxonomy club: Aloes (Asphodelaceae - subfamily Alooideae). At: Royal Botanic Gardens, Kew. London, UK. 28 February 2020.
Anthoons B, Madesis P, Drouzas A, de Boer H, 2020. Metabarcoding to trace adulteration in local Greek products. At: University of Ioannina. Ioannina, Greece. 31 January 2020.
Mascarello M, Beeckman H, Smets E, Janssens S, 2019. The use of high-throughput sequencing for the identification of illegally logged African trees. At: 5th Annual Meeting on Plant Ecology and Evolution. Meise Botanic Garden, Meise, Belgium. 29 November 2019.
Anthoons B, Madesis P, Drouzas A, de Boer H, 2019. Metabarcoding to trace adulteration in local Greek products. At: Cyprus University of Technology. Limassol, Cyprus. 22 November 2019.
Woudstra Y, 2020. Aloe identification improved - the quest for new DNA barcodes. At: Royal Horticultural Society. London, UK. 19 November 2019.
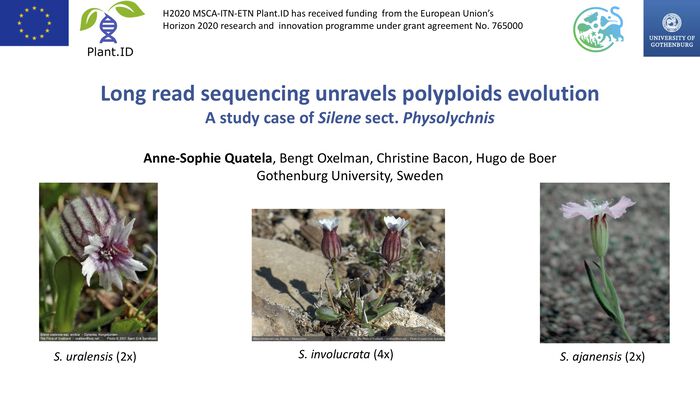
Quatela A-S, Oxelman B, Bacon C, de Boer H, 2019. Long read sequencing unravels polyploids evolution. At: Systematic Days. Gothenburg, Sweden. 18-19 November 2019.
Jahanbanifard M, Gravendeel B, Lens F, Verbeek FJ, 2019. Ebony wood identification to battle illegal trade. Biodiversity Information Science and Standards 3: e37084. http://doi.org/10.3897/biss.3.37084
Abstract also on Zenodo: https://zenodo.org/record/3251321#.XkagVXVKhwE
Polling M, de Boer HJ, Donders T, Verbeek FJ, Gravendeel B, 2019. Automatic pollen species image identification. At: Biodiversity_Next. Leiden, The Netherlands. 22-25 October 2019.
Canales NA, Walker K, 2019. Reconectando las colecciones de Cinchona en Madrid y Kew (Real expedición botánica al verreinato del Perú). At: Royal Botanical Garden Madrid. Madrid, Spain. October 2019.
Çiftçi O, Mertens A, van de Vijver B, Duijsings D, van Bodegom P, Gravendeel B, 2019. Transcriptomics for species delimitation in widespread bioindicator diatom, Nitzschia palea. At: Meeting of the Dutch-Flemish Society of Diatomists. Meise Botanic Garden, Belgium. 3-5 October 2019.
Cangren P, Jafari F, Moiloa N, Quatela A-S, Bacon C, Oxelman B, 2019. How long reads sequencing technologies enable reaching a double challenge in polyploid complexes: species delimitation by target capture from old herbarium specimens. At: Botany 2019. Tucson, Arizona, USA. 27-31 July 2019.
Anthoons B, Madesis P, Drouzas A, de Boer H, 2019. Metabarcoding to trace adulteration in local Greek products. At: 8th International Barcode of Life Conference. Trondheim, Norway. 17-20 June 2019.